Abstract
The origin of snake venom involved duplication and recruitment of non-venom genes into venom systems. Several studies have predicted that directional positive selection has governed this process. Venom composition varies substantially across snake species and venom phenotypes are locally adapted to prey, leading to coevolutionary interactions between predator and prey. Venom origins and contemporary snake venom evolution may therefore be driven by fundamentally different selection regimes, yet investigations of population-level patterns of selection have been limited. Here, we use whole-genome data from 68 rattlesnakes to test hypotheses about the factors that drive genomic diversity and differentiation in major venom gene regions. We show that selection has resulted in long-term maintenance of genetic diversity within and between species in multiple venom gene families. Our findings are inconsistent with a dominant role of directional positive selection and instead support a role of long-term balancing selection in shaping venom evolution. We also detect rapid decay of linkage disequilibrium due to high recombination rates in venom regions, suggesting that venom genes have reduced selective interference with nearby loci, including other venom paralogues. Our results provide an example of long-term balancing selection that drives trans-species polymorphism and help to explain how snake venom keeps pace with prey resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The genomic data that support the findings of this study are available at NCBI SRA under accession PRJNA593834.
Code availability
Analysis scripts are available on GitHub (https://github.com/drewschield/venom_population_genomics).
References
Zancolli, G. & Casewell, N. R. Venom systems as models for studying the origin and regulation of evolutionary novelties. Mol. Biol. Evol. 37, 2777–2790 (2020).
Arbuckle, K. From molecules to macroevolution: venom as a model system for evolutionary biology across levels of life. Toxicon X 6, 100034 (2020).
Mackessy, S. P. Handbook of Venoms and Toxins of Reptiles (CRC Press, 2021).
Hargreaves, A. D., Swain, M. T., Hegarty, M. J., Logan, D. W. & Mulley, J. F. Restriction and recruitment—gene duplication and the origin and evolution of snake venom toxins. Genome Biol. Evol. 6, 2088–2095 (2014).
Casewell, N. R., Huttley, G. A. & Wuster, W. Dynamic evolution of venom proteins in squamate reptiles. Nat. Commun. 3, 1066 (2012).
Casewell, N. R., Wuster, W., Vonk, F. J., Harrison, R. A. & Fry, B. G. Complex cocktails: the evolutionary novelty of venoms. Trends Ecol. Evol. 28, 219–229 (2013).
Fry, B. G. & Wuster, W. Assembling an arsenal: origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences. Mol. Biol. Evol. 21, 870–883 (2004).
Reyes-Velasco, J. et al. Expression of venom gene homologs in diverse python tissues suggests a new model for the evolution of snake venom. Mol. Biol. Evol. 32, 173–183 (2015).
Mackessy, S. P. in The Biology of Rattlesnakes (eds Hayes, W. K. et al.) 495–510 (Loma Linda Univ. Press, 2008).
Ikeda, N. et al. Unique structural characteristics and evolution of a cluster of venom phospholipase A 2 isozyme genes of Protobothrops flavoviridis snake. Gene 461, 15–25 (2010).
Dowell, N. L. et al. The deep origin and recent loss of venom toxin genes in rattlesnakes. Curr. Biol. 26, 2434–2445 (2016).
Schield, D. R. et al. The origins and evolution of chromosomes, dosage compensation, and mechanisms underlying venom regulation in snakes. Genome Res. 29, 590–601 (2019).
Lynch, V. J. Inventing an arsenal: adaptive evolution and neofunctionalization of snake venom phospholipase A2 genes. BMC Evol. Biol. 7, 2 (2007).
Casewell, N. R., Wagstaff, S. C., Harrison, R. A., Renjifo, C. & Wüster, W. Domain loss facilitates accelerated evolution and neofunctionalization of duplicate snake venom metalloproteinase toxin genes. Mol. Biol. Evol. 28, 2637–2649 (2011).
Mayr, E. Cause and effect in biology. Science 134, 1501–1506 (1961).
Aird, S. D. et al. Population genomic analysis of a pitviper reveals microevolutionary forces underlying venom chemistry. Genome Biol. Evol. 9, 2640–2649 (2017).
Margres, M. J. et al. Tipping the scales: the migration–selection balance leans toward selection in snake venoms. Mol. Biol. Evol. 36, 271–282 (2019).
Rautsaw, R. M. et al. Intraspecific sequence and gene expression variation contribute little to venom diversity in sidewinder rattlesnakes (Crotalus cerastes). Proc. R. Soc. B 286, 20190810 (2019).
Holding, M. L., Biardi, J. E. & Gibbs, H. L. Coevolution of venom function and venom resistance in a rattlesnake predator and its squirrel prey. Proc. R. Soc. B 283, 20152841 (2016).
Davies, E.-L. & Arbuckle, K. Coevolution of snake venom toxic activities and diet: evidence that ecological generalism favours toxicological diversity. Toxins 11, 711 (2019).
Smiley-Walters, S. A., Farrell, T. M. & Gibbs, H. L. Evaluating local adaptation of a complex phenotype: reciprocal tests of pigmy rattlesnake venoms on treefrog prey. Oecologia 184, 739–748 (2017).
Holding, M. L., Drabeck, D. H., Jansa, S. A. & Gibbs, H. L. Venom resistance as a model for understanding the molecular basis of complex coevolutionary adaptations. Integr. Comp. Biol. 56, 1032–1043 (2016).
Poran, N. S., Coss, R. G. & Benjamini, E. L. I. Resistance of California ground squirrels (Spermophilus beecheyi) to the venom of the northern Pacific rattlesnake (Crotalus viridis oreganus): a study of adaptive variation. Toxicon 25, 767–777 (1987).
Heatwole, H. & Poran, N. S. Resistances of sympatric and allopatric eels to sea snake venoms. Copeia 1995, 136–147 (1995).
Pomento, A. M., Perry, B. W., Denton, R. D., Gibbs, H. L. & Holding, M. L. No safety in the trees: local and species-level adaptation of an arboreal squirrel to the venom of sympatric rattlesnakes. Toxicon 118, 149–155 (2016).
Biardi, J. E., Chien, D. C. & Coss, R. G. California ground squirrel (Spermophilus beecheyi) defenses against rattlesnake venom digestive and hemostatic toxins. J. Chem. Ecol. 32, 137–154 (2006).
Jansa, S. A. & Voss, R. S. Adaptive evolution of the venom-targeted vWF protein in opossums that eat pitvipers. PLoS ONE 6, e20997 (2011).
Voss, R. S. & Jansa, S. A. Snake‐venom resistance as a mammalian trophic adaptation: lessons from didelphid marsupials. Biol. Rev. 87, 822–837 (2012).
Drabeck, D. H., Dean, A. M. & Jansa, S. A. Why the honey badger don’t care: convergent evolution of venom-targeted nicotinic acetylcholine receptors in mammals that survive venomous snake bites. Toxicon 99, 68–72 (2015).
Gibbs, H. L. et al. The molecular basis of venom resistance in a rattlesnake–squirrel predator–prey system. Mol. Ecol. 29, 2871–2888 (2020).
Kordiš, D., Bdolah, A. & Gubenšek, F. Positive Darwinian selection in Vipera palaestinae phospholipase A2 genes is unexpectedly limited to the third exon. Biochem. Biophys. Res. Commun. 251, 613–619 (1998).
Kordiš, D. & Gubenšek, F. Adaptive evolution of animal toxin multigene families. Gene 261, 43–52 (2000).
Juárez, P., Comas, I., González-Candelas, F. & Calvete, J. J. Evolution of snake venom disintegrins by positive Darwinian selection. Mol. Biol. Evol. 25, 2391–2407 (2008).
McDonald, J. H. & Kreitman, M. Adaptive protein evolution at the Adh locus in Drosophila. Nature 351, 652–654 (1991).
Strickland, J. L. et al. Evidence for divergent patterns of local selection driving venom variation in Mojave Rattlesnakes (Crotalus scutulatus). Sci. Rep. 8, 17622 (2018).
Przeworski, M., Coop, G. & Wall, J. D. The signature of positive selection on standing genetic variation. Evolution 59, 2312–2323 (2005).
Cutter, A. D. & Payseur, B. A. Genomic signatures of selection at linked sites: unifying the disparity among species. Nat. Rev. Genet. 14, 262 (2013).
Aquadro, C. F., Begun, D. J. & Kindahl, E. C. in Non-Neutral Evolution (ed. Golding, B.) 46–56 (Springer, 1994).
Begun, D. J. & Aquadro, C. F. Molecular population genetics of the distal portion of the X chromosome in Drosophila: evidence for genetic hitchhiking of the yellow-achaete region. Genetics 129, 1147–1158 (1991).
Maynard Smith, J. & Haigh, J. The hitch-hiking effect of a favourable gene. Genet. Res. 23, 23–35 (1974).
Barton, N. H. Genetic hitchhiking. Philos. Trans. R. Soc. Lond. B 355, 1553–1562 (2000).
Charlesworth, D. Balancing selection and its effects on sequences in nearby genome regions. PLoS Genet. 2, e64 (2006).
Fijarczyk, A. & Babik, W. Detecting balancing selection in genomes: limits and prospects. Mol. Ecol. 24, 3529–3545 (2015).
Piertney, S. B. & Oliver, M. K. The evolutionary ecology of the major histocompatibility complex. Heredity 96, 7–21 (2006).
Kelley, J., Walter, L. & Trowsdale, J. Comparative genomics of major histocompatibility complexes. Immunogenetics 56, 683–695 (2005).
Bakker, E. G., Toomajian, C., Kreitman, M. & Bergelson, J. A genome-wide survey of R gene polymorphisms in Arabidopsis. Plant Cell 18, 1803–1818 (2006).
Goldberg, E. E. et al. Species selection maintains self-incompatibility. Science 330, 493–495 (2010).
Llaurens, V., Whibley, A. & Joron, M. Genetic architecture and balancing selection: the life and death of differentiated variants. Mol. Ecol. 26, 2430–2448 (2017).
Thompson, J. N. The Geographic Mosaic of Coevolution (Univ. Chicago Press, 2005).
Yoder, J. B. & Nuismer, S. L. When does coevolution promote diversification? Am. Nat. 176, 802–817 (2010).
Leffler, E. M. et al. Multiple instances of ancient balancing selection shared between humans and chimpanzees. Science 339, 1578–1582 (2013).
DeGiorgio, M., Lohmueller, K. E. & Nielsen, R. A model-based approach for identifying signatures of ancient balancing selection in genetic data. PLoS Genet. 10, e1004561 (2014).
Takahata, N. A simple genealogical structure of strongly balanced allelic lines and trans-species evolution of polymorphism. Proc. Natl Acad. Sci. USA 87, 2419–2423 (1990).
Clark, A. G. Neutral behavior of shared polymorphism. Proc. Natl Acad. Sci. USA 94, 7730–7734 (1997).
Wiuf, C., Zhao, K., Innan, H. & Nordborg, M. The probability and chromosomal extent of trans-specific polymorphism. Genetics 168, 2363–2372 (2004).
Teixeira, J. C. et al. Long-term balancing selection in LAD1 maintains a missense trans-species polymorphism in humans, chimpanzees, and bonobos. Mol. Biol. Evol. 32, 1186–1196 (2015).
Hill, W. G. & Robertson, A. The effect of linkage on limits to artificial selection. Genet. Res. 8, 269–294 (1966).
Barton, N. H. & Charlesworth, B. Why sex and recombination? Science 281, 1986–1990 (1998).
Webster, M. T. & Hurst, L. D. Direct and indirect consequences of meiotic recombination: implications for genome evolution. Trends Genet. 28, 101–109 (2012).
McGaugh, S. E. et al. Recombination modulates how selection affects linked sites in Drosophila. PLoS Biol. 10, e1001422 (2012).
Begun, D. J. & Aquadro, C. F. Levels of naturally occurring DNA polymorphism correlate with recombination rates in D. melanogaster. Nature 356, 519–520 (1992).
Schield, D. R. et al. Snake recombination landscapes are directed by PRDM9 but concentrated in functional regions. Mol. Biol. Evol. 37, 1272–1294 (2020).
Mackessy, S. P. Evolutionary trends in venom composition in the Western Rattlesnakes (Crotalus viridis sensu lato): toxicity vs. tenderizers. Toxicon 55, 1463–1474 (2010).
Holding, M. L., Sovic, M. G., Colston, T. J. & Gibbs, H. L. The scales of coevolution: comparative phylogeography and genetic demography of a locally adapted venomous predator and its prey. Biol. J. Linn. Soc. 132, 297–317 (2021).
Schield, D. R. et al. Allopatric divergence and secondary contact with gene flow: a recurring theme in rattlesnake speciation. Biol. J. Linn. Soc. 128, 149–169 (2019).
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Sabeti, P. C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Ferrer-Admetlla, A., Liang, M., Korneliussen, T. & Nielsen, R. On detecting incomplete soft or hard selective sweeps using haplotype structure. Mol. Biol. Evol. 31, 1275–1291 (2014).
Gao, Z., Przeworski, M. & Sella, G. Footprints of ancient‐balanced polymorphisms in genetic variation data from closely related species. Evolution 69, 431–446 (2015).
Siewert, K. M. & Voight, B. F. Detecting long-term balancing selection using allele frequency correlation. Mol. Biol. Evol. 34, 2996–3005 (2017).
Siewert, K. M. & Voight, B. F. BetaScan2: standardized statistics to detect balancing selection utilizing substitution data. Genome Biol. Evol. 12, 3873–3877 (2020).
Cheng, X. & DeGiorgio, M. Flexible mixture model approaches that accommodate footprint size variability for robust detection of balancing selection. Mol. Biol. Evol. 37, 3267–3291 (2020).
Cheng, X. & DeGiorgio, M. BalLeRMix+: mixture model approaches for robust joint identification of both positive selection and long-term balancing selection. Bioinformatics 38, 861–863 (2022).
Navarro, A. & Barton, N. H. The effects of multilocus balancing selection on neutral variability. Genetics 161, 849–863 (2002).
Fry, B. G. From genome to ‘venome’: molecular origin and evolution of the snake venom proteome inferred from phylogenetic analysis of toxin sequences and related body proteins. Genome Res. 15, 403–420 (2005).
Bernardoni, J. L. et al. Functional variability of snake venom metalloproteinases: adaptive advantages in targeting different prey and implications for human envenomation. PLoS ONE 9, e109651 (2014).
Modahl, C. M., Mrinalini, Frietze, S. & Mackessy, S. P. Adaptive evolution of distinct prey-specific toxin genes in rear-fanged snake venom. Proc. R. Soc. B 285, 20181003 (2018).
Klauber, L. M. Rattlesnakes: Their Habits, Life Histories, and Influence on Mankind (Univ. California Press, 1956).
Holding, M. L. et al. Phylogenetically diverse diets favor more complex venoms in North American pitvipers. Proc. Natl Acad. Sci. USA 118, e2015579118 (2021).
Axel, B., Pook, C. E., Harrison, R. A. & Wolfgang, W. Coevolution of diet and prey-specific venom activity supports the role of selection in snake venom evolution. Proc. R. Soc. B 276, 2443–2449 (2009).
Margres, M. J. et al. The Tiger Rattlesnake genome reveals a complex genotype underlying a simple venom phenotype. Proc. Natl Acad. Sci. USA 118, e2014634118 (2021).
Mason, A. J. et al. Trait differentiation and modular toxin expression in palm-pitvipers. BMC Genomics 21, 147 (2020).
Clarke, B. C. The evolution of genetic diversity. Proc. R. Soc. B. 205, 453–474 (1979).
Arbuckle, K., de la Vega, R. C. R. & Casewell, N. R. Coevolution takes the sting out of it: evolutionary biology and mechanisms of toxin resistance in animals. Toxicon 140, 118–131 (2017).
Schield, D. R., Perry, B. W., Nikolakis, Z. L., Mackessy, S. P. & Castoe, T. A. Population genomic analyses confirm male-biased mutation rates in snakes. J. Hered. 112, 221–227 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Van der Auwera, G. A. et al. From FastQ data to high‐confidence variant calls: the genome analysis toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 10–11 (2013).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Delaneau, O., Howie, B., Cox, A. J., Zagury, J.-F. & Marchini, J. Haplotype estimation using sequencing reads. Am. J. Hum. Genet. 93, 687–696 (2013).
Dowell, N. L. et al. Extremely divergent haplotypes in two toxin gene complexes encode alternative venom types within rattlesnake species. Curr. Biol. 28, 1016–1026 (2018).
Xie, C. & Tammi, M. T. CNV-seq, a new method to detect copy number variation using high-throughput sequencing. BMC Bioinf. 10, 80 (2009).
Wigginton, J. E., Cutler, D. J. & Abecasis, G. R. A note on exact tests of Hardy–Weinberg equilibrium. Am. J. Hum. Genet. 76, 887–893 (2005).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Green, R. E. et al. Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs. Science 346, 1254449 (2014).
Korunes, K. L. & Samuk, K. pixy: unbiased estimation of nucleotide diversity and divergence in the presence of missing data. Mol. Ecol. Resour. 21, 1359–1368 (2021).
Nei, M. & Roychoudhury, A. K. Sampling variances of heterozygosity and genetic distance. Genetics 76, 379–390 (1974).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Gautier, M. & Vitalis, R. rehh: an R package to detect footprints of selection in genome-wide SNP data from haplotype structure. Bioinformatics 28, 1176–1177 (2012).
Gautier, M., Klassmann, A. & Vitalis, R. rehh 2.0: a reimplementation of the R package rehh to detect positive selection from haplotype structure. Mol. Ecol. Resour. 17, 78–90 (2017).
Renaud, G. glactools: a command-line toolset for the management of genotype likelihoods and allele counts. Bioinformatics 34, 1398–1400 (2018).
Haller, B. C. & Messer, P. W. SLiM 3: forward genetic simulations beyond the Wright–Fisher model. Mol. Biol. Evol. 36, 632–637 (2019).
Chan, A. H., Jenkins, P. A. & Song, Y. S. Genome-wide fine-scale recombination rate variation in Drosophila melanogaster. PLoS Genet. 8, e1003090 (2012).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2017).
Acknowledgements
We thank R. Orton and N. Balchan for assistance in the field. We thank J. Vindum and the California Academy of Sciences for tissue loans. A. Ludington kindly provided advice on demographic analysis. We thank S. Flaxman and R. Safran for helpful discussion on the study and feedback on the manuscript. This work was supported by National Science Foundation (NSF) postdoctoral research fellowship grant DBI-1906188 to D.R.S., NSF grant DEB-1501886 to D.R.S. and T.A.C. and NSF grant DEB-1655571 to T.A.C., S.P.M. and J.M.M., NSF grants DEB-1949268, BCS-2001063 and DBI-2130666 to M.D. and National Institutes of Health grant R35GM128590 to M.D.
Author information
Authors and Affiliations
Contributions
D.R.S. and T.A.C. designed the study. D.R.S., B.W.P., Z.L.N., S.S.G., C.F.S., J.M.P., J.M.M., S.P.M. and T.A.C. collected samples and generated data. D.R.S., B.W.P., R.H.A. and M.D. performed analyses. D.R.S. and T.A.C. wrote the manuscript with contributions from B.W.P., R.H.A., M.L.H. and M.D. All authors provided edits to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Giulia Zancolli and Wieslaw Babik for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Overview of filtering strategy to remove potential bioinformatic artifacts of copy-number variation (CNV) in major venom gene regions.
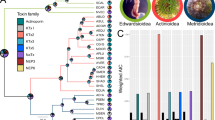
a Schematic representation of the workflow used to filter departures from Hardy–Weinberg equilibrium (HWE) due to excess heterozygosity at SNP positions. Input variant calls were analysed using a one-tailed HWE test (adapted from Wigginton et al. 2005) and SNPs with significant HWE P-values after FDR correction were filtered. Individual CNVs were detected, scored, and regional variation in CNV presence, quantified as the proportion of individuals in a population with a detected CNV (% CNV), was measured across each major venom gene region. CNV calls were used to mask genotypes per individual prior to population genetic analysis. b Regional variation in CNV presence per population across the major venom gene regions. Top panels for each venom gene region show log2 read depths in sliding windows relative to the autosomal median depth for the CV1 reference population. Here values of zero indicate equal coverage to the autosomal median, values of −1 equal half coverage, and values of 1 equal twice the autosomal median coverage. Dark blue segments indicate the locations of venom genes in each region. Lower panels for each region show variation in the proportion of individuals with masked genotypes in a detected CNV (% CNV) per population. c Minor allele frequency spectra across the dataset for each gene in the major venom gene families after using the described HWE and CNV filtering strategy.
Extended Data Fig. 2 Genome scans of genetic diversity within populations and differentiation between populations across chromosome housing major venom gene families (SVMPs, SVSPs, and PLA2s) in CV2 and CO2 populations.
a Sliding windows of nucleotide diversity (π) in C. viridis (CV2) and C. oreganus (CO2) populations and sequence divergence (dxy) and relative differentiation (Fst) between CV1 and CV2 and between CO1 and CO2 across Chromosome 9 (top panels), and in the SVMP region (bottom panels). b π, dxy, and Fst across Chromosome 10 (top), and in the SVSP region (bottom). c π, dxy, and Fst across Chromosome 15 (top), and in the PLA2 region (bottom). Shaded points in top panels show estimates in 10 kb windows and lines show estimates in 100 kb windows. In bottom panels in a and b, shaded points show estimates in 1 kb windows, and lines show estimates in 10 kb windows. In bottom panels in c, shaded points show estimates in 250 bp windows and lines are estimates in 1 kb windows. The regions housing venom genes are shaded in grey in all panels. Chromosome-specific and genome-wide mean values for each statistic are represented by blue and red dashed horizontal lines. The locations of individual venom genes are shown as blue boxes (bottom panels). The non-venom homologue PLA2gIIE is shown in light purple. Gaps in measurements are locations that were masked due to significant evidence of copy-number variation between C. viridis and C. oreganus.
Extended Data Fig. 3 Genetic diversity point estimates for major venom paralogues.
Point estimates (dark blue circles) of π (a), dxy (b), and Fst (c) for paralogues in the three major venom gene families, compared to chromosome-specific background distributions outside of venom gene regions. Boxplots show the median (horizontal lines), interquartile (box limits), and range (whiskers) based on n = 2,253, 1,998, and 1,239 10 kb sliding windows for chromosomes 9, 10, and 15, respectively.
Extended Data Fig. 4 Tajima’s D across major venom regions in CV2 and CO2 populations.
Tajima’s D across SVMP (a), SVSP (b), and PLA2 (c) regions in CV2 and CO2 populations. Top panels show sliding window Tajima’s D estimates in 100 kb (lines) and 10 kb (points points) windows. Venom gene regions are shaded in grey. Middle panels in a and b show zoomed in venom gene regions with sliding window estimates in 10 kb (lines) and 1 kb (points) windows. Middle panels in c show estimates in 1 kb (lines) and 250 bp (points) windows. Gaps in lines represent windows where there was insufficient data to calculate a mean estimate. Dark blue segments show the locations of venom genes in each region, and the non-venom homologue PLA2gIIE is shown in light purple in c. Regional variation in the presence of CNVs (% CNV) in CV2 (orange dashed line) and CO2 (dark blue line) is shown below venom region scans. Individual genotypes in detected CNVs were masked. Bottom panels show distributions of chromosome-specific and non-venom homologue (NV) backgrounds compared to values in each venom gene region, with boxplots showing the median (horizontal lines), interquartile (box limits), and range (whiskers). Asterisks indicate significant differences between venom gene regions and chromosome backgrounds and non-venom homologues based on two-tailed Welch’s two-sample t-tests and n = 2,293, 2,068, and 1,272 10 kb sliding windows for SVMP, SVSP, and PLA2 comparisons, respectively (*P < 0.05; **P < 0.001). Exact P-values for comparisons can be found in Supplementary Table 5.
Extended Data Fig. 5 Genomic scans of iHS and ß selection statistics in CV1 and CO1 populations.
Lines show mean estimates in 1 Mb sliding windows. Shaded points show mean estimates in 100 kb sliding windows.
Extended Data Fig. 6 Scans of iHS and ß selection statistics across major venom regions.
Scans of a|iHS|and b ß selection statistics in CV1 and CO1 populations across SVMP, SVSP, and PLA2 venom gene regions. Lines in SVMP and SVSP panels show mean estimates in 10 kb sliding windows. Shaded points in SVMP and SVSP panels show mean estimates in 1 kb sliding windows. Lines in PLA2 panels show mean estimates in 1 kb sliding windows. Shaded points in PLA2 panels show mean estimates in 250 bp sliding windows. Gaps in lines represent windows where there was insufficient data to calculate a mean estimate. Venom gene locations are shown with dark blue segments. The non-venom PLA2gIIE homologue is shown as a light purple segment.
Extended Data Fig. 7 Selection statistic point estimates for major venom paralogues.
Point estimates (dark blue circles) of Tajima’s D (a), df (b), |iHS|(c), and ß (d) for paralogues in the three major venom gene families, compared to chromosome-specific background distributions outside of venom gene regions. Boxplots show the median (horizontal lines), interquartile (box limits), and range (whiskers) based on n = 2,253, 1,998, and 1,239 10 kb sliding windows for chromosomes 9, 10, and 15, respectively.
Extended Data Fig. 8 Diagnostic parameters in composite-likelihood ratio tests of selection in major venom regions.
Results of composite-likelihood ratio tests of balancing selection and diagnostic parameters in the SVMP (a-b), SVSP (c-d), and PLA2 (e-f) venom gene regions. Top and middle panels show B0,MAF scores, log\(\widehat {(A)}\), and \(\hat x\) parameters as shown in Fig. 3 and Extended Data Fig. 9. Dark blue arrows show locations of venom genes and dashed lines show the genome-wide 95th quantile. Lower panels show the dispersion parameter, log\(\widehat {(a)}\), where positive values indicate balancing selection in regions with high B0,MAF scores and negative values are indicative of positive selection. The dashed horizontal line indicates 0 on the y-axis.
Extended Data Fig. 9 Signatures of selection in the PLA2 venom gene region.
Signatures of selection in the PLA2 venom gene region, with comparisons to chromosomal backgrounds and non-venom homologues (NV). a Tajima’s D across chromosome 15 in C. viridis (CV1) and C. oreganus (CO1) populations (top). Middle panels show variation zoomed into the PLA2 region, shaded in grey. Boxplots show distributions of Tajima’s D for chromosome 15, non-venom homologues of the PLA2 family, and PLA2s. b Proportion of fixed differences (df) between CV1 and CO1. c Integrated haplotype statistics (|iHS|) in CV1 and CO1. d ß statistic measuring allele frequency correlation in CV1 and CO1. Points in chromosome scan panels represent mean estimates in 10 kb sliding windows, and lines represent 100 kb windowed estimates. Points in zoomed PLA2 region scans represent mean estimates in 250 bp windows and lines show 1 kb windowed estimates. Locations of individual PLA2 genes are shown as dark blue segments (bottom panels). The PLA2gIIE non-venom homologue is shown in light purple. Gaps in lines represent windows where there was insufficient data to calculate a mean estimate. Boxplots in a-d show the median (horizontal lines), interquartile (box limits), and range (whiskers). Asterisks indicate significant differences between venom gene regions and chromosome backgrounds and non-venom homologues based on two-tailed Welch’s two-sample t-tests (Tajima’s D and |iHS|) and Mann–Whitney ∪ tests (df and ß) and n = 1,272 10 kb sliding windows for comparisons (*P < 0.05; **P < 0.001; ***P < 2.2 × 10−16). Exact P-values for comparisons can be found in Supplementary Table 5. e-f Results of composite-likelihood ratio tests of balancing selection versus neutrality using B0,MAF scores in the PLA2 region in CV1 and CO1. Upper panels show B0,MAF scores, with higher values indicating greater evidence for balancing selection. Arrows show locations of PLA2 genes. Lower panels show inferred footprint size, log\(\widehat {(A)}\), as solid grey lines and equilibrium allele frequency, \(\hat x\), as dashed lines. Dashed lines in top panels of e-f show the genome-wide 95th quantile.
Extended Data Fig. 10 Population-scaled recombination rates in major venom regions.
Population-scaled recombination rate (ρ = 4Ner) across SVMP (a), SVSP (b), and PLA2 (c) venom gene regions in C. viridis and C. oreganus. Upper panels show chromosome-wide variation and lower panels show variation within the venom regions, specifically, highlighted by the grey shading in all panels. Dark blue segments in lower panels show the locations of venom paralogues. The light purple segment in c is the non-venom homologue PLA2gIIE. Shaded points in the upper panels of a-c represent mean ρ in 10 kb windows and black lines represent 100 kb windowed means. In lower panels of a and b, points and lines represent 1 kb and 10 kb windowed ρ. In lower panels of c, lines represent 1 kb windowed ρ and points are estimates from all SNP intervals. Vertical dashed lines show the locations of inferred recombination hotspots from Schield et al. (2020). Panels at the bottom show regional variation in the proportion of C. viridis (orange dashed line) and C. oreganus (dark blue line) individuals per population with evidence of copy-number variation (% CNV) across the venom gene regions.
Supplementary information
Supplementary Information
Supplementary Results, Discussion, Methods, References, Figs. 1–5 and Tables 1–11.
Rights and permissions
About this article
Cite this article
Schield, D.R., Perry, B.W., Adams, R.H. et al. The roles of balancing selection and recombination in the evolution of rattlesnake venom. Nat Ecol Evol 6, 1367–1380 (2022). https://doi.org/10.1038/s41559-022-01829-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-022-01829-5
This article is cited by
-
Pangenomics of the death cap mushroom Amanita phalloides, and of Agaricales, reveals dynamic evolution of toxin genes in an invasive range
The ISME Journal (2023)
-
Sequence Divergence in Venom Genes Within and Between Montane Pitviper (Viperidae: Crotalinae: Cerrophidion) Species is Driven by Mutation–Drift Equilibrium
Journal of Molecular Evolution (2023)